PPE: Last Millennium Reanalysis#
[1]:
%load_ext autoreload
%autoreload 2
import os
os.chdir('/glade/work/fengzhu/GitHub/cfr/docsrc/v2026/notebooks')
import pandas as pd
import xarray as xr
import xesmf as xe
import numpy as np
import cfr
cfr.use('v2026')
print(cfr.__version__)
2026.4.8
Pseudoproxies Setup#
[23]:
df_raw = pd.read_json('https://github.com/fzhu2e/cfr-data/raw/refs/heads/main/pages2kv2.json')
df = df_raw.rename(columns={
'paleoData_pages2kID': 'pid',
'geo_meanLat': 'lat',
'geo_meanLon': 'lon',
'archiveType': 'archive',
})[['pid', 'lat', 'lon', 'archive']]
# mask = df['archive'].isin(['marine sediment', 'coral', 'sclerosponge', 'bivalve'])
# obs = cfr.Obs(df[mask])
obs = cfr.Obs(df)
obs.df['time'] = object
obs.df['value'] = object
[2]:
dirpath = '/glade/work/fengzhu/Data/GCM_sims'
ds_prior = xr.open_dataset(os.path.join(dirpath, 'CCSM4', 'tas_sfc_Amon_CCSM4_past1000_085001-185012.nc'))
ds_truth = xr.open_dataset(os.path.join(dirpath, 'iCESM', 'LM', 'tas_sfc_Amon_iCESM_past1000historical_085001-200512.nc'))
ds_truth_ann = ds_truth.x.annualize(time2year=True) - 273.15
ds_truth_ann['tas'].attrs['units'] = 'degC'
ds_prior = ds_prior.drop_vars(['lat_bnds', 'lon_bnds', 'height'])
ds_prior_ann = ds_prior.x.annualize(time2year=True) - 273.15
ds_prior_ann['tas'].attrs['units'] = 'degC'
regridder = xe.Regridder(ds_prior_ann, ds_truth_ann, method='bilinear')
ds_prior_ann = regridder(ds_prior_ann)
ds_prior_ann['tas'].attrs = ds_truth_ann['tas'].attrs
[24]:
# a simple white noise model without data attrition
SNR = 10 # var(signal) / var(noise)
for idx, row in obs.df.iterrows():
np.random.seed(idx)
obs.df.at[idx, 'time'] = np.arange(850, 2006)
obs.df.at[idx, 'SNR'] = SNR
signal = ds_truth_ann.sel(
lat=row['lat'],
lon=row['lon'],
method='nearest',
).tas.values
signal_var = np.var(signal, ddof=1)
noise_var = signal_var / SNR
obs.df.at[idx, 'R'] = noise_var
noise = np.random.normal(loc=0, scale=np.sqrt(noise_var), size=signal.shape)
obs.df.at[idx, 'value'] = signal + noise
obs.df['type'] = 'T'
obs.df['psm_name'] = 'IdenticalTS'
obs.df
[24]:
| pid | lat | lon | archive | time | value | SNR | R | type | psm_name | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NAm_153 | 52.70 | 241.70 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [-0.02292751859725406, 0.3572480420356854, 0.7... | 10.0 | 0.089662 | T | IdenticalTS |
| 1 | Asi_245 | 23.00 | 114.00 | documents | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [21.125607818090135, 20.116943803724595, 19.96... | 10.0 | 0.021568 | T | IdenticalTS |
| 2 | NAm_165 | 37.90 | 252.30 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [6.338173780630901, 7.4904591259960585, 6.8232... | 10.0 | 0.065712 | T | IdenticalTS |
| 3 | Asi_178 | 28.77 | 83.73 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [7.1370074286774425, 5.966696792272471, 6.3234... | 10.0 | 0.046389 | T | IdenticalTS |
| 4 | Asi_174 | 28.18 | 85.43 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [10.911502987956526, 10.585710012500336, 10.76... | 10.0 | 0.013729 | T | IdenticalTS |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 687 | Asi_201 | 35.88 | 74.18 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [-4.287775723319058, -4.232198470857572, -3.74... | 10.0 | 0.047043 | T | IdenticalTS |
| 688 | Asi_179 | 27.50 | 88.02 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [7.563148014053656, 7.277179376837475, 7.47118... | 10.0 | 0.019030 | T | IdenticalTS |
| 689 | Arc_014 | 63.62 | 29.10 | lake sediment | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [-1.0157051133439026, -0.25941108796600443, 1.... | 10.0 | 0.212980 | T | IdenticalTS |
| 690 | Ocn_071 | 16.20 | 298.51 | coral | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [25.283580513311232, 24.776707927084082, 24.91... | 10.0 | 0.012692 | T | IdenticalTS |
| 691 | Ocn_072 | 16.20 | 298.51 | coral | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [25.144609174862826, 24.651241206523522, 24.64... | 10.0 | 0.012692 | T | IdenticalTS |
692 rows × 10 columns
[25]:
np.unique(obs.df['archive'])
[25]:
array(['bivalve', 'borehole', 'coral', 'documents', 'glacier ice',
'hybrid', 'lake sediment', 'marine sediment', 'sclerosponge',
'speleothem', 'tree'], dtype=object)
[26]:
os.makedirs('./data', exist_ok=True)
obs.df.to_json('./data/df_proxy_ocn.json')
df_ocn = pd.read_json('./data/df_proxy_ocn.json')
print(type(df_ocn.loc[0, 'time']))
print(type(df_ocn.loc[0, 'value']))
df_ocn
<class 'list'>
<class 'list'>
[26]:
| pid | lat | lon | archive | time | value | SNR | R | type | psm_name | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NAm_153 | 52.70 | 241.70 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [-0.022927518600000002, 0.35724804200000004, 0... | 10 | 0.089662 | T | IdenticalTS |
| 1 | Asi_245 | 23.00 | 114.00 | documents | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [21.1256078181, 20.1169438037, 19.9622574808, ... | 10 | 0.021568 | T | IdenticalTS |
| 2 | NAm_165 | 37.90 | 252.30 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [6.3381737806, 7.490459126, 6.8232782039, 7.80... | 10 | 0.065712 | T | IdenticalTS |
| 3 | Asi_178 | 28.77 | 83.73 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [7.1370074287, 5.9666967923000005, 6.323457105... | 10 | 0.046389 | T | IdenticalTS |
| 4 | Asi_174 | 28.18 | 85.43 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [10.911502988, 10.5857100125, 10.7688062264, 1... | 10 | 0.013729 | T | IdenticalTS |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 687 | Asi_201 | 35.88 | 74.18 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [-4.2877757233, -4.2321984709, -3.7413174658, ... | 10 | 0.047043 | T | IdenticalTS |
| 688 | Asi_179 | 27.50 | 88.02 | tree | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [7.5631480141, 7.2771793768, 7.471187646, 7.01... | 10 | 0.019030 | T | IdenticalTS |
| 689 | Arc_014 | 63.62 | 29.10 | lake sediment | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [-1.0157051133, -0.259411088, 1.4312935975, -2... | 10 | 0.212980 | T | IdenticalTS |
| 690 | Ocn_071 | 16.20 | 298.51 | coral | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [25.2835805133, 24.7767079271, 24.9194705363, ... | 10 | 0.012692 | T | IdenticalTS |
| 691 | Ocn_072 | 16.20 | 298.51 | coral | [850, 851, 852, 853, 854, 855, 856, 857, 858, ... | [25.1446091749, 24.6512412065, 24.6425996453, ... | 10 | 0.012692 | T | IdenticalTS |
692 rows × 10 columns
[27]:
obs.setup()
obs['Ocn_065'].data
[27]:
pid Ocn_065
lat 25.84
lon 281.38
archive coral
time [850, 851, 852, 853, 854, 855, 856, 857, 858, ...
value [24.868690830872385, 24.688057548488253, 24.62...
SNR 10.0
R 0.006686
type T
psm_name IdenticalTS
Name: 12, dtype: object
[28]:
cfr.set_style('journal', font_scale=1.2)
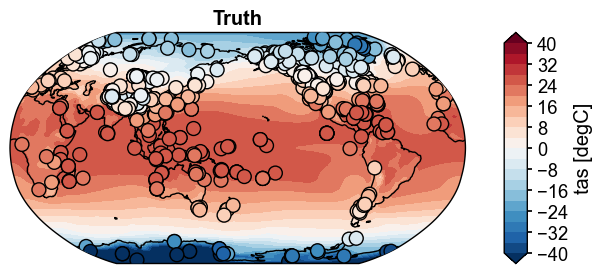
fig, ax = obs.plot(
levels=np.linspace(-40, 40, 21),
cbar_kwargs={'ticks': np.linspace(-40, 40, 11)},
cyclic=True,
da_target=ds_truth_ann['tas'],
t_idx=-1,
title='Truth',
)
Prior Setup#
[29]:
pm = cfr.PriorMember(ds_prior_ann)
pm.gen_samples_bootstrap(nens=30, clim_yrs=1)
prior = cfr.Prior(pm)
prior.ds
[29]:
<xarray.Dataset> Size: 2MB
Dimensions: (lat: 96, lon: 144, ens: 30)
Coordinates:
* lat (lat) float32 384B -90.0 -88.11 -86.21 -84.32 ... 86.21 88.11 90.0
* lon (lon) float32 576B 0.0 2.5 5.0 7.5 10.0 ... 350.0 352.5 355.0 357.5
Dimensions without coordinates: ens
Data variables:
tas (lat, lon, ens) float32 2MB -47.45 -47.06 -48.57 ... -23.29 -20.72[30]:
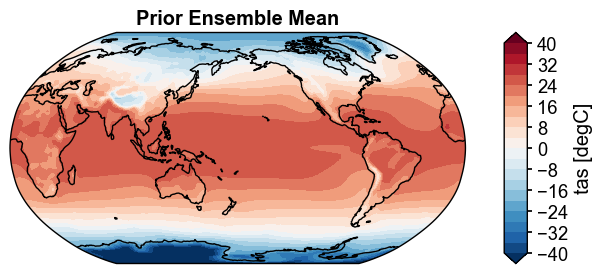
fig, ax = prior.ds['tas'].mean('ens').x.plot(
levels=np.linspace(-40, 40, 21),
cbar_kwargs={'ticks': np.linspace(-40, 40, 11)},
cyclic=True,
title='Prior Ensemble Mean',
)
EnKF Solver Setup#
[31]:
prior.ds_rgd = prior.ds.copy()
prior.ds_ann = prior.ds.copy()
da_solver = cfr.enkf.Solver(prior, obs)
da_solver.prep(
recon_period=(1801, 1850),
loc_radius=10000,
)
>>> Proxy System Modeling: Y = H(X)
>>> Computing the localization matrix
Processing variables: 100%|██████████| 1/1 [00:00<00:00, 1.90it/s]
[32]:
# da_solver.run(nproc=1)
[33]:
%%time
da_solver.run(nproc=4, chunksize=10)
>>> DA update
Updating time slices: 100%|██████████| 50/50 [00:12<00:00, 4.06it/s]
CPU times: user 1.33 s, sys: 2.22 s, total: 3.55 s
Wall time: 12.8 s
[34]:
os.makedirs('./recons', exist_ok=True)
da_solver.post.to_netcdf(f'./recons/recon_LMR_SNR10_ocn_loc10000.nc')
da_solver.post
[34]:
<xarray.Dataset> Size: 166MB
Dimensions: (time: 50, lat: 96, lon: 144, ens: 30)
Coordinates:
* lat (lat) float32 384B -90.0 -88.11 -86.21 -84.32 ... 86.21 88.11 90.0
* lon (lon) float32 576B 0.0 2.5 5.0 7.5 10.0 ... 350.0 352.5 355.0 357.5
* time (time) int64 400B 1801 1802 1803 1804 1805 ... 1847 1848 1849 1850
Dimensions without coordinates: ens
Data variables:
tas (time, lat, lon, ens) float64 166MB -53.43 -53.49 ... -23.44 -23.22Validation#
[35]:
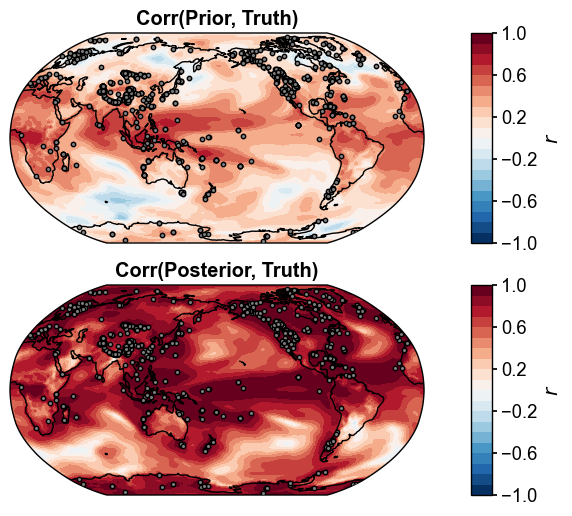
fig, ax = da_solver.plot_valid(
ds_truth_ann, ds_prior=ds_prior_ann, vn='tas', metric='corr',
df_sites=obs.df[['lat', 'lon']],
site_markersizes=10,
legend=False,
cyclic=True,
)
[36]:
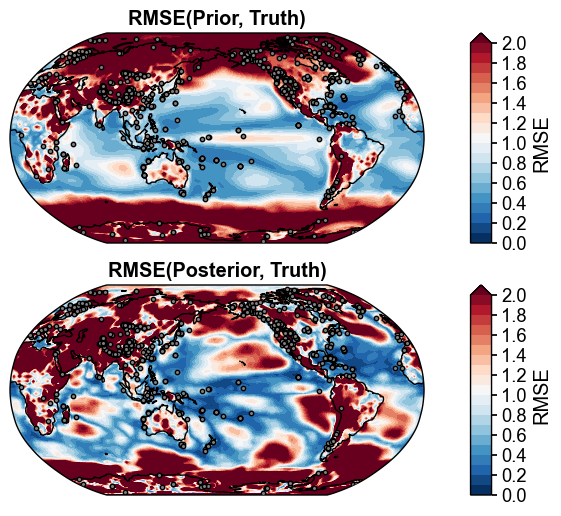
fig, ax = da_solver.plot_valid(
ds_truth_ann, ds_prior=ds_prior_ann, vn='tas', metric='RMSE',
df_sites=obs.df[['lat', 'lon']],
site_markersizes=10,
legend=False,
cyclic=True,
)
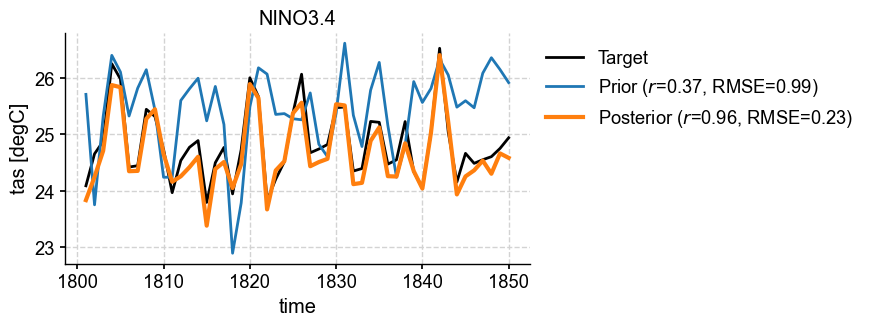
[37]:
fig, ax = da_solver.plot_valid(ds_truth_ann, ds_prior=ds_prior_ann, metric='nino3.4')
[ ]:
